OBJECTIVE:
To lay down the procedure for Identification and Isolation of Microorganisms.
SCOPE:
This SOP is applicable for the procedure for Identification and Isolation of Microorganisms at {Company Name} {Location}.
RESPONSIBILITY:
Microbiologist: is responsible to perform the activity as per SOP.
In charge- Microbiology- is responsible to ensure compliance as per SOP.
Head/Designee Quality Control – Shall be responsible for ensuring compliance as per SOP.
ACCOUNTABILITY:
QA Head shall be Accountable for implementation of SOP.
ABOUT IDENTIFICATION OF MICRO ORGANISM:
The identification and isolation of microorganisms are fundamental processes in microbiology. Isolation techniques, such as streak plating, spread plating, and pour plating, aim to obtain pure cultures by separating individual cells on a growth medium.
Once isolated, microorganisms are identified through a combination of methods. These include microscopic examination (cell shape, size, and arrangement), staining procedures (like Gram staining), biochemical tests (to determine metabolic capabilities), and molecular techniques (such as DNA sequencing). Accurate identification is crucial in various fields, including medicine, research, and industry.
PROCEDURE:
Precaution:
Streak isolated colonies to be identified in such a manner to get an isolated colony.
Pick up isolated colonies only.
Follow aseptic practices to avoid any contamination.
Identification criteria:
Identification up to species level in case of counts observed in grade ‘A’ area.
Identification up to species level in case microbial counts exceeds alert/ action/specification limit in purified water, environmental monitoring in grade ‘C’ and ‘D’ areas.
Identification up to species level in case of OOT and OOS results.
Identification up to species level in case of growth having typical/atypical characteristics observed on selective agar.
Identification up to species level whenever new colonies are observed.
Isolation of Organisms:
Prepare the media as per the SOP.
At the end of the incubation period of Environmental monitoring, Water Analysis and MLT analysis examine the plates for suspected colonies.
If suspected colonies are found proceed for isolation.
Pick up the suspected colony.Streak the colony on 48 hours pre incubated Soya bean Casein Digest Agar medium plates.
Incubate the plates for 18-24 hours at 30-35° C.
In case of fungal isolates incubate the plate at 22.5±2.5°C for 5 days.
Select the isolated colony and find out the colony characteristics and cell arrangement as follows and enter the details as per Format-I.
Various colony edges:
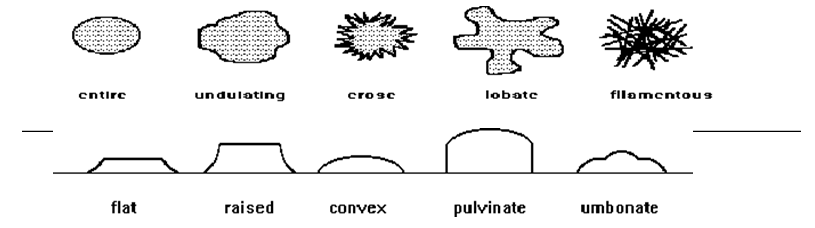
Cell arrangement:
Coccus (plural, cocci): Cells those are spherical in shape. The picture below shows cocci arranged singly.
Streptococcus: A bacterium with spherical cells arranged in chains, like beads on a string.
Staphylococcus: A bacterium with spherical cells arranged in clusters, like clusters of grapes.
Diplococcus: A bacterium with spherical cells arranged in pairs.
Tetrad: Spherical bacterial cells arranged in a group of four. A tetrad arrangement looks almost like a re-square under the microscope.
Sarcina (plural, Sarcinae): Spherical bacterial cells arranged in a group of eight. Sarcinae look like small cubes and may be difficult to distinguish from tetrads.
Bacillus (plural, bacilli): A bacterium with rod-shaped cells. The picture below shows Bacilli arranged singly.
Diplobacillus: A bacterium with rod-shaped cells arranged in pairs.
Streptobacillus: A bacterium with rod-shaped cells arranged in end-to-end chains. Streptobacilli often resemble link sausages.
Coryneform bacillus: A bacterium with irregularly rod-shaped cells arranged at angles to form V- and L-shaped arrangements.
Spirillum (plural, spirilla): A bacterium with cells that are rigid and spiral in shape.
Vibrio: A bacterium with curved or comma-shaped cells.
Spirochete: A bacterium with flexible, spiral-shaped cells. Spirochetes often appear helical or corkscrew-shaped with tapered ends.
Cell arrangements:
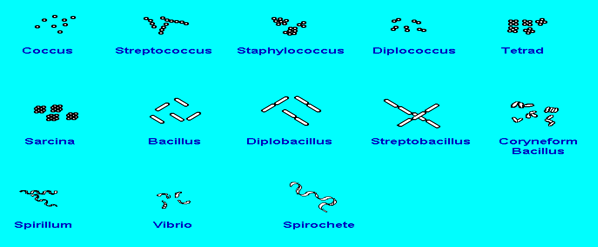
Isolation of standard microorganisms:
Whenever new ATCC cultures are received or during the preparation of primary and working cultures, identify the bacterial isolate up to genus level and species level by out sourcing the test. In case of fungi obtained identify the isolate up to genus level
Isolation of microorganism on rotational basis:
If all of the above criteria for identification do not allow to identifying the organisms (eg: No count is observed in grade ‘A’, monitoring samples are within alert/action/specification limits), select organisms randomly and identify them up to species level at least once in six months.
Isolation from utility samples and environmental monitoring (from grades ‘A’, ‘C’ and ‘D’):
Isolate the microorganism, when the microbial counts exceed alert, action and specification limits or when the morphologically different colonies are observed. Identify them up to species level and create a database for the cultures obtained in Format-II.
Update the data base as and when a new isolate is identified.
Identify the predominant and new flora when the total microbial counts exceed alert, action and specification limits.
The characterization of the predominant and new flora shall be useful in investigating the excursions.
Due to the physiological stress on the isolates, the organisms inherently may show up slightly different morphology.
In this context, if an isolate is found morphologically different, the isolate shall be picked and identified.
In case this organism is already identified (repeat isolate), then the database shall be updated accordingly, but the culture need not be preserved.
Verify the growth promoting ability of the media using the isolate initially.Identify the bacterial isolate up to genus level and species level by out sourcing the test.
In case of fungi obtained from monitoring, identify the isolate up to genus level.
The identified isolates shall be preserved as primary cultures. Prepare the required number of primary cultures as per SOP entitled.
Identification of bacterial isolates: Conduct the orientation checks like gram staining, spore staining, motility, shape, color, elevation, margin, surface, arrangement and biochemical test for the bacterial isolates.
Gram staining: Perform gram staining as per the GTP.
Spore staining: Perform spore staining as per the GTP.
Motility test:
Prepare the suspension and take a loopful of culture and mix with saline.Clean and flame the hanging drop slide (Depression slide) and place it on the table with the depression uppermost.
Spread a little Vaseline or petroleum jelly around the cavity of the slide.
Clean the cover slip and apply petroleum jelly on each of four corners of the cover slip, using matchstick or stick.
Place the cover slip on clean paper with petroleum jelly up.
Place the loopful of suspension in the center of the cover slip.
Place the depression slide on the cover slip, with the cavity facing down so that the depression covers the suspension.
Press the slide gently to form a seal between the cover slip and the slide.
Quickly turn the cavity slide so that cover slip should be at upper side end suspension should be hanging inside the cavity.
Take care while turning the slide so that suspension in hanging drop manner.
Examine the preparation under low power objective with reduced light and also observe the slide under high power and oil immersion objective.
Record the observations.Inoculate an isolated colony of the organism to be tested using a needle into motility media.
Click the link to download word file copy of this document:
https://pharmaguidehub.com/product/identification-and-isolation-of-microorganisms/
Simply stab the media in a straight line as possible and withdraw the needle very carefully to avoid destroying the straight line.
Incubate the tube at 32.5 ± 2.5°C for 24 to 48 hours.
After incubation check whether the bacteria have migrated away from the original line of inoculation
Results: Migration of bacteria away from the line of inoculation indicates a positive test; lack of migration away from the line of inoculation indicates a negative test.
Catalase test:
Take a small amount of the test organism onto the slide.
Add a drop of 3% v/v Hydrogen peroxide to the smear.
Result: An instant evolution of visible bubbles indicates positive catalase reaction, while lack of evolution of bubbles indicates absence of catalase.
Oxidase test:
Take the Oxidase disc in a sterile plastic petri dish and add the test organism with the help of inoculation loop.
Observe the disc for colour change.
Result: The appearance of a dark purple/ blue colour for 30 to 60 seconds indicates a positive reaction, while absence of colour indicates negatives.
Click the link to download word file copy of this document:
https://pharmaguidehub.com/product/identification-and-isolation-of-microorganisms/
Indole test:
Inoculate an isolated colony of the organism to be tested in 5 mL of peptone water (pure culture) and incubate for 24 ± 2 hours at 32.5 ± 2.5°C.
After incubation add 0.2 to 0.3 mL of Kovac’s Indole reagent, shake gently and leave it to stand for about 10 minutes.
Result: A dark red/purple colour in the reagent layer indicates a positive reaction. No change in original colour of the reagent indicates negative test.
Methyl red test:
Inoculate an isolated colony of the organism to be tested in 5 mL of MR-VP medium (Glucose phosphate broth) and incubate for 24 ± 2 hours at 32.5 ± 2.5°C.
Add about 5 drops of Methyl-red indicator, mix the medium and observe the colour change.
Result: A bright red colour indicates positive reaction and yellow colour indicates negative test.
Voges proskauer test:
Inoculate an isolated colony of the organism to be tested in 5 mL of MR-VP medium (Glucose phosphate broth) and incubate for 24 ± 2 hours at 32.5 ± 2.5°C.
To 1 mL of culture add 0.6 mL of 5% w/v a-napthal solution and 0.2 mL of 40% w/v Potassium Hydroxide solution.
Shake well and observe the broth after 15 minutes for the colour change.
Result: Development of a pink to crimson colour within 15 minutes constitutes a positive test and no change in colour indicates negative test.
Citrate utilization test:
Streak an isolated colony of the organism to be tested in Simmon’s citrate agar slant, do not stab the butt.
Incubate for 24 to 48 hours at 32.5 ± 2.5°C with a loose cap.
Result: A prussian blue color slant indicates positive test while negative test will have no bacterial growth and will remain green in colour
Urease test:
Inoculate a loopful of fresh bacterial culture over the entire slope surface of Christenson’s agar medium and incubate for 24 to 48 hours at 32.5 ± 2.5°C.
After incubation observe the medium for any change in colour.
Colour of the medium should change from yellow to purple pink indicating positive reaction for urease test, while lack of colour change indicates negative reaction.
Hydrogen sulphide production test:
Inoculate an isolated colony of the test organism on the slant of Triple sugar iron agar by stabbing the butt using an inoculation needle and streaking the slope using an inoculation loop.
Incubate the tube for 24 to 48 hours at 32.5 ± 2.5°C.
After incubation observe the tube for blackening of the butt due to Hydrogen sulphide production.
Sugar fermentation test:
Filter sterilize specific carbohydrates such as Glucose, Mannitol, Maltose, Lactose, Sucrose Dulcitol and Xylose at a concentration of 1% separately and add aseptically into phenol red broth and mark all the tubes as 1, 2, 3…etc.
Inoculate a loopful of test culture in all the tubes.
Incubate the tubes at 32.5 ± 2.5°C for 48 hours and observe for acid production.
Record all the results in Format-I.
Ammonia production test:
Prepare peptone water.
Inoculate the culture and incubated at 37± 2°C for 48 hrs.
At the end of incubation, add Nesslers reagent. Brown color indicates positive test.
Coagulase test:
Coagulase is a soluble enzyme like a product that converts the fibrinogen in citrated human or rabbit plasma into fibrin aided by an activator in the plasma.
Reconstitute the rabbit plasma by adding sterile high pure water as per the volume mentioned on the product label.
Mix the entire content by gentle rotation of the tube and allow it to dissolve completely.
Add a drop of young culture to 0.5 mL of rabbit plasma and incubate at 37±2°C for 1 hour.
A positive test is shown by clotting of plasma.
Tabulate the results of all the tests performed as above.
Fungal Staining:
Place a drop of lacto phenol cotton blue on a clean glass slide.
Transfer small tuft of fungus, preferably with spores and spore bearing structures.Mix gently stain with mould structures.
Place the cover glass over the preparation taking care to avoid air bubble in the stain and observe the stain under microscope initially at 10x and then under 45x.
Allotment of number to isolates:
Each environmental isolates shall be assigned with a unique in-house identification number.
The identification number shall consist of seven characters.
First two letters represent the source of microorganisms.
Next two numeric letters denote the year.
Next three numeric characters indicate serial number of the isolate starting from 001 and increasing by 1 for next isolate.
For example, EM13001 indicates first environmental isolate of the year 2013.
Abbreviation of source for allotment of isolate number:
EM: Environmental monitoring
UT: Utility samples PS: Product samples
Maintenance of isolates:
Take a coloured photograph showing colony characteristics of identified isolate for colony morphology when it is identified for first time and maintain the photographic library.
The photograph of identified isolates shall be assigned with a unique in-house identification number as PG/xx/xxxx.
First two letters represent photograph.
Next two numeric letters denote the year.
Next four numeric characters indicate serial number of the photograph starting from 0001 and increasing by 1 for next isolate.
For example, PG280001 indicates first photograph of the identified isolate for the year 2028.
Use this library for matching colony morphology of subsequent microorganism identification.
Maintain the culture in primary cultures as per the SOP.
Prepare the working cultures of environmental isolates from primary cultures as and when required.
Prepare the culture suspension from working cultures as per the SOP as and when required.
The isolates shall be preserved up to preparation of annual summary report.Based on annual summary report, minimum six numbers of in-house floras shall be selected and preserved based on the predominance and type of activity.
These micro-floras shall be maintained as “In house flora” for one year period.
Out of six in-house floras per activity, the first predominant four in-house floras shall be used in routine microbiological studies and later two shall be kept as a “reserved isolates”.
During course of maintenance and usage, if any of the four in-house floras are unable to grow, it shall be discontinued and replaced with reserved isolate.
Other isolates shall be decontaminated at 121°C for 30 minutes as per SOP.
Click the link to download word file copy of this document:
https://pharmaguidehub.com/product/identification-and-isolation-of-microorganisms/
Annual summary Report and Data trending:
Summary report of identification shall be prepared once in a year containing including
List of all isolates identified throughout the year based on the activity.
Pre-dominancy of specific flora and its source based on the type of activity.
Trending of typical flora on percent basis.
Correlation of isolated flora to alert/action results in case of monitoring results if any.
Correlation of isolated flora to OOT/OOS results in case of raw material and finished product results, if any.
Recommendation for usage of predominant microbial flora in routine microbiology studies based on the source from where they are isolated.
REFERENCES:
Not Applicable
ANNEXURES:
| ANNEXURE NO. | TITLE OF ANNEXURE |
| Annexure-I | Identification of microorganisms |
| Annexure-II | Culture Library |
ENCLOSURES: SOP Training Record.
DISTRIBUTION:
- Controlled Copy No. 01 : Head Quality Assurance
- Controlled Copy No. 02 : Head Quality Control (Micro.)
- Master Copy : Quality Assurance Department
ABBREVIATIONS:
| No. | : | Number |
| °C | : | Degree Centigrade |
| OOS | : | Out of specification |
| WC | : | Working Culture |
| OOT | : | Out of trend |
| SOP | : | Standard Operating Procedure |
| QC | : | Quality Control |
REVISION HISTORY:
CHANGE HISTORY LOG
| Revision No. | Details of Changes | Reason for Change | Effective Date |
| 00 | New SOP | Not Applicable | To Be Written Manual |
Click the link to download word file copy of this document:
https://pharmaguidehub.com/product/identification-and-isolation-of-microorganisms/
Annexure-I
IDENTIFICATION OF MICROORGANISMS


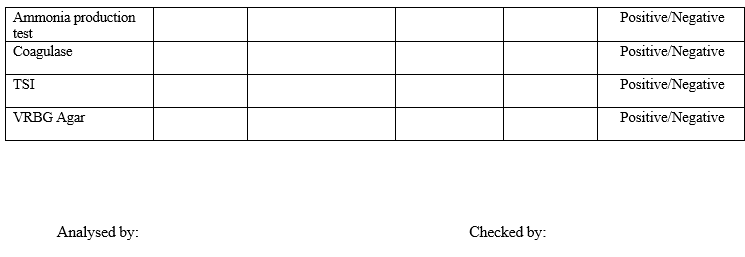
Annexure-II
CULTURE LIBRARY
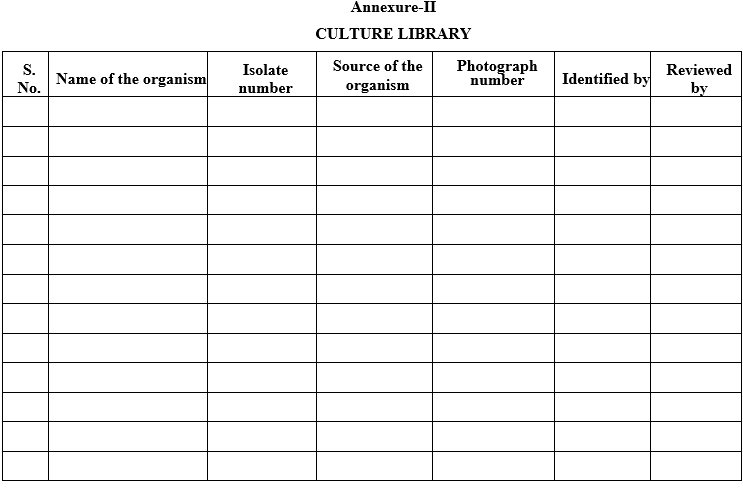
Click the link to download word file copy of this document:
https://pharmaguidehub.com/product/identification-and-isolation-of-microorganisms/

